Protein backbone angles1/16/2024 
Besides being the parts of the main chain, each amino acid, starting from its C α atom, has a side chain as well. Typically ω is fixed at 180 ∘ for majority proteins 4, and so only ϕ and ψ are to be determined. 
1, protein backbone structures can essentially be represented by dihedral angles ϕ, ψ, and ω, which are respectively defined by taking every four consecutive atoms from the sequence C i - 1, N i, C α i, C i, N i + 1, C α i. 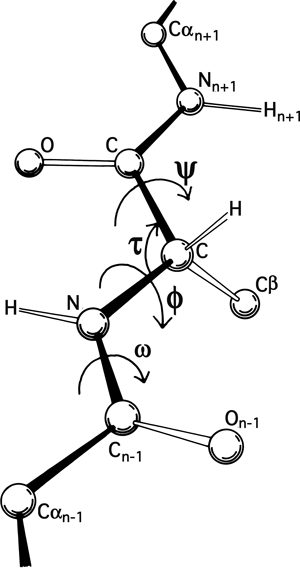
The C and N atoms of every two consecutive amino acids in a protein form a peptide bond and thus we obtain the backbone or main chain of the protein. Each amino acid has three common atoms N, C α and C among others. A protein might have any of the 20 types of amino acids any number of times in any order subject to stoichiometric constraints 3. The difficulties come from the inevitability of searching an astronomically large conformation space and from the absence of a highly accurate energy function to evaluate potential protein conformations 2. The PSP problem is to determine the three dimensional structures of given proteins just from their amino acid sequences. Three dimensional structures of most proteins depend on their amino acid (AA) sequences. Protein structure prediction (PSP) has been an unsolved problem for the last half century 1. The SAP program along with its data is available from the website. We then empirically show that SAP significantly outperforms existing state-of-the-art methods on well-known benchmark datasets: for some types of angles, the differences are above 3 in mean absolute error (MAE). With these arguments in mind, we present a deep learning method named Simpler Angle Predictor (SAP) to train simpler DNN models that enhance protein backbone angle prediction. We also argue that comparatively simpler predictors can more easily be reconstructed than the more complex ones. From artificial intelligence and machine learning perspectives, problem representations and solution approaches do mutually interact and thus affect performance. While more features might add more predictive power to the neural network, we argue that redundant features could rather clutter the scenario and more complex neural networks then just could counterbalance the noise. However, we observe that in the protein backbone angle prediction research, there is an overall trend to employ more and more complex neural networks and then to throw more and more features to the neural networks. Prediction of protein structures via the representations using backbone dihedral angles has recently achieved significant progress along with the on-going surge of deep neural network (DNN) research in general. Protein structure prediction is a grand challenge.
0 Comments
Leave a Reply.AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
